W3cubDocs
/scikit-imageskimage.color
Color space conversion.
Stain to RGB color space conversion. | |
Convert an image array to a new color space. | |
Euclidean distance between two points in Lab color space | |
Color difference as given by the CIEDE 2000 standard. | |
Color difference according to CIEDE 94 standard | |
Color difference from the CMC l:c standard. | |
Create an RGB representation of a gray-level image. | |
Create a RGBA representation of a gray-level image. | |
Haematoxylin-Eosin-DAB (HED) to RGB color space conversion. | |
HSV to RGB color space conversion. | |
Convert image in CIE-LAB to CIE-LCh color space. | |
Convert image in CIE-LAB to sRGB color space. | |
Convert image in CIE-LAB to XYZ color space. | |
Return an RGB image where color-coded labels are painted over the image. | |
Convert image in CIE-LCh to CIE-LAB color space. | |
Luv to RGB color space conversion. | |
CIE-Luv to XYZ color space conversion. | |
Compute luminance of an RGB image. | |
RGB to Haematoxylin-Eosin-DAB (HED) color space conversion. | |
RGB to HSV color space conversion. | |
Conversion from the sRGB color space (IEC 61966-2-1:1999) to the CIE Lab colorspace under the given illuminant and observer. | |
RGB to CIE-Luv color space conversion. | |
RGB to RGB CIE color space conversion. | |
RGB to XYZ color space conversion. | |
RGB to YCbCr color space conversion. | |
RGB to YDbDr color space conversion. | |
RGB to YIQ color space conversion. | |
RGB to YPbPr color space conversion. | |
RGB to YUV color space conversion. | |
RGBA to RGB conversion using alpha blending [1]. | |
RGB CIE to RGB color space conversion. | |
RGB to stain color space conversion. | |
XYZ to CIE-LAB color space conversion. | |
XYZ to CIE-Luv color space conversion. | |
XYZ to RGB color space conversion. | |
Get the CIE XYZ tristimulus values. | |
YCbCr to RGB color space conversion. | |
YDbDr to RGB color space conversion. | |
YIQ to RGB color space conversion. | |
YPbPr to RGB color space conversion. | |
YUV to RGB color space conversion. |
-
skimage.color.combine_stains(stains, conv_matrix, *, channel_axis=-1)[source] -
Stain to RGB color space conversion.
- Parameters:
-
-
stains(…, C=3, …) array_like -
The image in stain color space. By default, the final dimension denotes channels.
- conv_matrix: ndarray
-
The stain separation matrix as described by G. Landini [1].
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in RGB format. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If
stainsis not at least 2-D with shape (…, C=3, …).
Notes
Stain combination matrices available in the
colormodule and their respective colorspace:-
rgb_from_hed: Hematoxylin + Eosin + DAB -
rgb_from_hdx: Hematoxylin + DAB -
rgb_from_fgx: Feulgen + Light Green -
rgb_from_bex: Giemsa stain : Methyl Blue + Eosin -
rgb_from_rbd: FastRed + FastBlue + DAB -
rgb_from_gdx: Methyl Green + DAB -
rgb_from_hax: Hematoxylin + AEC -
rgb_from_bro: Blue matrix Anilline Blue + Red matrix Azocarmine + Orange matrix Orange-G -
rgb_from_bpx: Methyl Blue + Ponceau Fuchsin -
rgb_from_ahx: Alcian Blue + Hematoxylin -
rgb_from_hpx: Hematoxylin + PAS
References
[1][2]A. C. Ruifrok and D. A. Johnston, “Quantification of histochemical staining by color deconvolution,” Anal. Quant. Cytol. Histol., vol. 23, no. 4, pp. 291–299, Aug. 2001.
Examples
>>> from skimage import data >>> from skimage.color import (separate_stains, combine_stains, ... hdx_from_rgb, rgb_from_hdx) >>> ihc = data.immunohistochemistry() >>> ihc_hdx = separate_stains(ihc, hdx_from_rgb) >>> ihc_rgb = combine_stains(ihc_hdx, rgb_from_hdx)
-
skimage.color.convert_colorspace(arr, fromspace, tospace, *, channel_axis=-1)[source] -
Convert an image array to a new color space.
- Valid color spaces are:
-
‘RGB’, ‘HSV’, ‘RGB CIE’, ‘XYZ’, ‘YUV’, ‘YIQ’, ‘YPbPr’, ‘YCbCr’, ‘YDbDr’
- Parameters:
-
-
arr(…, C=3, …) array_like -
The image to convert. By default, the final dimension denotes channels.
-
fromspacestr -
The color space to convert from. Can be specified in lower case.
-
tospacestr -
The color space to convert to. Can be specified in lower case.
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The converted image. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If fromspace is not a valid color space
- ValueError
-
If tospace is not a valid color space
Notes
Conversion is performed through the “central” RGB color space, i.e. conversion from XYZ to HSV is implemented as
XYZ -> RGB -> HSVinstead of directly.Examples
>>> from skimage import data >>> img = data.astronaut() >>> img_hsv = convert_colorspace(img, 'RGB', 'HSV')
-
skimage.color.deltaE_cie76(lab1, lab2, channel_axis=-1)[source] -
Euclidean distance between two points in Lab color space
- Parameters:
-
-
lab1array_like -
reference color (Lab colorspace)
-
lab2array_like -
comparison color (Lab colorspace)
-
channel_axisint, optional -
This parameter indicates which axis of the arrays corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
dEarray_like -
distance between colors
lab1andlab2
-
References
[2]A. R. Robertson, “The CIE 1976 color-difference formulae,” Color Res. Appl. 2, 7-11 (1977).
-
skimage.color.deltaE_ciede2000(lab1, lab2, kL=1, kC=1, kH=1, *, channel_axis=-1)[source] -
Color difference as given by the CIEDE 2000 standard.
CIEDE 2000 is a major revision of CIDE94. The perceptual calibration is largely based on experience with automotive paint on smooth surfaces.
- Parameters:
-
-
lab1array_like -
reference color (Lab colorspace)
-
lab2array_like -
comparison color (Lab colorspace)
-
kLfloat (range), optional -
lightness scale factor, 1 for “acceptably close”; 2 for “imperceptible” see deltaE_cmc
-
kCfloat (range), optional -
chroma scale factor, usually 1
-
kHfloat (range), optional -
hue scale factor, usually 1
-
channel_axisint, optional -
This parameter indicates which axis of the arrays corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
deltaEarray_like -
The distance between
lab1andlab2
-
Notes
CIEDE 2000 assumes parametric weighting factors for the lightness, chroma, and hue (
kL,kC,kHrespectively). These default to 1.References
[3]M. Melgosa, J. Quesada, and E. Hita, “Uniformity of some recent color metrics tested with an accurate color-difference tolerance dataset,” Appl. Opt. 33, 8069-8077 (1994).
-
skimage.color.deltaE_ciede94(lab1, lab2, kH=1, kC=1, kL=1, k1=0.045, k2=0.015, *, channel_axis=-1)[source] -
Color difference according to CIEDE 94 standard
Accommodates perceptual non-uniformities through the use of application specific scale factors (
kH,kC,kL,k1, andk2).- Parameters:
-
-
lab1array_like -
reference color (Lab colorspace)
-
lab2array_like -
comparison color (Lab colorspace)
-
kHfloat, optional -
Hue scale
-
kCfloat, optional -
Chroma scale
-
kLfloat, optional -
Lightness scale
-
k1float, optional -
first scale parameter
-
k2float, optional -
second scale parameter
-
channel_axisint, optional -
This parameter indicates which axis of the arrays corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
dEarray_like -
color difference between
lab1andlab2
-
Notes
deltaE_ciede94 is not symmetric with respect to lab1 and lab2. CIEDE94 defines the scales for the lightness, hue, and chroma in terms of the first color. Consequently, the first color should be regarded as the “reference” color.
kL,k1,k2depend on the application and default to the values suggested for graphic artsParameter
Graphic Arts
Textiles
kL1.000
2.000
k10.045
0.048
k20.015
0.014
References
-
skimage.color.deltaE_cmc(lab1, lab2, kL=1, kC=1, *, channel_axis=-1)[source] -
Color difference from the CMC l:c standard.
This color difference was developed by the Colour Measurement Committee (CMC) of the Society of Dyers and Colourists (United Kingdom). It is intended for use in the textile industry.
The scale factors
kL,kCset the weight given to differences in lightness and chroma relative to differences in hue. The usual values arekL=2,kC=1for “acceptability” andkL=1,kC=1for “imperceptibility”. Colors withdE > 1are “different” for the given scale factors.- Parameters:
-
-
lab1array_like -
reference color (Lab colorspace)
-
lab2array_like -
comparison color (Lab colorspace)
-
channel_axisint, optional -
This parameter indicates which axis of the arrays corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
dEarray_like -
distance between colors
lab1andlab2
-
Notes
deltaE_cmc the defines the scales for the lightness, hue, and chroma in terms of the first color. Consequently
deltaE_cmc(lab1, lab2) != deltaE_cmc(lab2, lab1)References
[3]F. J. J. Clarke, R. McDonald, and B. Rigg, “Modification to the JPC79 colour-difference formula,” J. Soc. Dyers Colour. 100, 128-132 (1984).
-
skimage.color.gray2rgb(image, *, channel_axis=-1)[source] -
Create an RGB representation of a gray-level image.
- Parameters:
-
-
imagearray_like -
Input image.
-
channel_axisint, optional -
This parameter indicates which axis of the output array will correspond to channels.
-
- Returns:
-
-
rgb(…, C=3, …) ndarray -
RGB image. A new dimension of length 3 is added to input image.
-
Notes
If the input is a 1-dimensional image of shape
(M,), the output will be shape(M, C=3).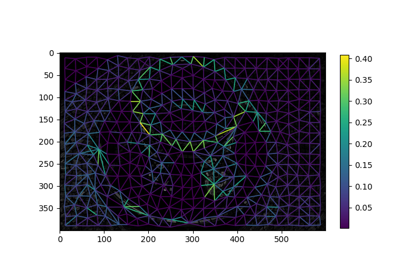
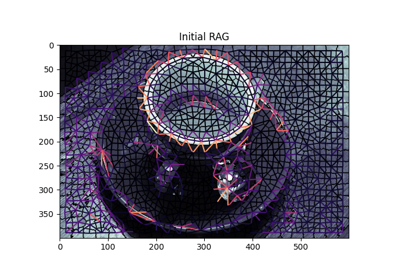
Region Boundary based Region adjacency graphs (RAGs)
Region Boundary based Region adjacency graphs (RAGs)
-
skimage.color.gray2rgba(image, alpha=None, *, channel_axis=-1)[source] -
Create a RGBA representation of a gray-level image.
- Parameters:
-
-
imagearray_like -
Input image.
-
alphaarray_like, optional -
Alpha channel of the output image. It may be a scalar or an array that can be broadcast to
image. If not specified it is set to the maximum limit corresponding to theimagedtype. -
channel_axisint, optional -
This parameter indicates which axis of the output array will correspond to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
rgbandarray -
RGBA image. A new dimension of length 4 is added to input image shape.
-
-
skimage.color.hed2rgb(hed, *, channel_axis=-1)[source] -
Haematoxylin-Eosin-DAB (HED) to RGB color space conversion.
- Parameters:
-
-
hed(…, C=3, …) array_like -
The image in the HED color space. By default, the final dimension denotes channels.
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in RGB. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If
hedis not at least 2-D with shape (…, C=3, …).
References
[1]A. C. Ruifrok and D. A. Johnston, “Quantification of histochemical staining by color deconvolution.,” Analytical and quantitative cytology and histology / the International Academy of Cytology [and] American Society of Cytology, vol. 23, no. 4, pp. 291-9, Aug. 2001.
Examples
>>> from skimage import data >>> from skimage.color import rgb2hed, hed2rgb >>> ihc = data.immunohistochemistry() >>> ihc_hed = rgb2hed(ihc) >>> ihc_rgb = hed2rgb(ihc_hed)
-
skimage.color.hsv2rgb(hsv, *, channel_axis=-1)[source] -
HSV to RGB color space conversion.
- Parameters:
-
-
hsv(…, C=3, …) array_like -
The image in HSV format. By default, the final dimension denotes channels.
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in RGB format. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If
hsvis not at least 2-D with shape (…, C=3, …).
Notes
Conversion between RGB and HSV color spaces results in some loss of precision, due to integer arithmetic and rounding [1].
References
Examples
>>> from skimage import data >>> img = data.astronaut() >>> img_hsv = rgb2hsv(img) >>> img_rgb = hsv2rgb(img_hsv)
-
skimage.color.lab2lch(lab, *, channel_axis=-1)[source] -
Convert image in CIE-LAB to CIE-LCh color space.
CIE-LCh is the cylindrical representation of the CIE-LAB (Cartesian) color space.
- Parameters:
-
-
lab(…, C=3, …) array_like -
The input image in CIE-LAB color space. Unless
channel_axisis set, the final dimension denotes the CIE-LAB channels. The L* values range from 0 to 100; the a* and b* values range from -128 to 127. -
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in CIE-LCh color space, of same shape as input.
-
- Raises:
-
- ValueError
-
If
labdoes not have at least 3 channels (i.e., L*, a*, and b*).
See also
Notes
The h channel (i.e., hue) is expressed as an angle in range
(0, 2*pi).References
Examples
>>> from skimage import data >>> from skimage.color import rgb2lab, lab2lch >>> img = data.astronaut() >>> img_lab = rgb2lab(img) >>> img_lch = lab2lch(img_lab)
-
skimage.color.lab2rgb(lab, illuminant='D65', observer='2', *, channel_axis=-1)[source] -
Convert image in CIE-LAB to sRGB color space.
- Parameters:
-
-
lab(…, C=3, …) array_like -
The input image in CIE-LAB color space. Unless
channel_axisis set, the final dimension denotes the CIE-LAB channels. The L* values range from 0 to 100; the a* and b* values range from -128 to 127. -
illuminant{“A”, “B”, “C”, “D50”, “D55”, “D65”, “D75”, “E”}, optional -
The name of the illuminant (the function is NOT case sensitive).
-
observer{“2”, “10”, “R”}, optional -
The aperture angle of the observer.
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in sRGB color space, of same shape as input.
-
- Raises:
-
- ValueError
-
If
labis not at least 2-D with shape (…, C=3, …).
See also
Notes
This function uses
lab2xyz()andxyz2rgb(). The CIE XYZ tristimulus values are x_ref = 95.047, y_ref = 100., and z_ref = 108.883. See functionxyz_tristimulus_values()for a list of supported illuminants.References
-
skimage.color.lab2xyz(lab, illuminant='D65', observer='2', *, channel_axis=-1)[source] -
Convert image in CIE-LAB to XYZ color space.
- Parameters:
-
-
lab(…, C=3, …) array_like -
The input image in CIE-LAB color space. Unless
channel_axisis set, the final dimension denotes the CIE-LAB channels. The L* values range from 0 to 100; the a* and b* values range from -128 to 127. -
illuminant{“A”, “B”, “C”, “D50”, “D55”, “D65”, “D75”, “E”}, optional -
The name of the illuminant (the function is NOT case sensitive).
-
observer{“2”, “10”, “R”}, optional -
The aperture angle of the observer.
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in XYZ color space, of same shape as input.
-
- Raises:
-
- ValueError
-
If
labis not at least 2-D with shape (…, C=3, …). - ValueError
-
If either the illuminant or the observer angle are not supported or unknown.
- UserWarning
-
If any of the pixels are invalid (Z < 0).
See also
Notes
The CIE XYZ tristimulus values are x_ref = 95.047, y_ref = 100., and z_ref = 108.883. See function
xyz_tristimulus_values()for a list of supported illuminants.References
-
skimage.color.label2rgb(label, image=None, colors=None, alpha=0.3, bg_label=0, bg_color=(0, 0, 0), image_alpha=1, kind='overlay', *, saturation=0, channel_axis=-1)[source] -
Return an RGB image where color-coded labels are painted over the image.
- Parameters:
-
-
labelndarray -
Integer array of labels with the same shape as
image. -
imagendarray, optional -
Image used as underlay for labels. It should have the same shape as
labels, optionally with an additional RGB (channels) axis. Ifimageis an RGB image, it is converted to grayscale before coloring. -
colorslist, optional -
List of colors. If the number of labels exceeds the number of colors, then the colors are cycled.
-
alphafloat [0, 1], optional -
Opacity of colorized labels. Ignored if image is
None. -
bg_labelint, optional -
Label that’s treated as the background. If
bg_labelis specified,bg_colorisNone, andkindisoverlay, background is not painted by any colors. -
bg_colorstr or array, optional -
Background color. Must be a name in
skimage.color.color_dictor RGB float values between [0, 1]. -
image_alphafloat [0, 1], optional -
Opacity of the image.
-
kindstring, one of {‘overlay’, ‘avg’} -
The kind of color image desired. ‘overlay’ cycles over defined colors and overlays the colored labels over the original image. ‘avg’ replaces each labeled segment with its average color, for a stained-class or pastel painting appearance.
-
saturationfloat [0, 1], optional -
Parameter to control the saturation applied to the original image between fully saturated (original RGB,
saturation=1) and fully unsaturated (grayscale,saturation=0). Only applies whenkind='overlay'. -
channel_axisint, optional -
This parameter indicates which axis of the output array will correspond to channels. If
imageis provided, this must also match the axis ofimagethat corresponds to channels.Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
resultndarray of float, same shape as image -
The result of blending a cycling colormap (
colors) for each distinct value inlabelwith the image, at a certain alpha value.
-
Use pixel graphs to find an object’s geodesic center
Use pixel graphs to find an object's geodesic centerComparing edge-based and region-based segmentation
Comparing edge-based and region-based segmentation
-
skimage.color.lch2lab(lch, *, channel_axis=-1)[source] -
Convert image in CIE-LCh to CIE-LAB color space.
CIE-LCh is the cylindrical representation of the CIE-LAB (Cartesian) color space.
- Parameters:
-
-
lch(…, C=3, …) array_like -
The input image in CIE-LCh color space. Unless
channel_axisis set, the final dimension denotes the CIE-LAB channels. The L* values range from 0 to 100; the C values range from 0 to 100; the h values range from 0 to2*pi. -
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in CIE-LAB format, of same shape as input.
-
- Raises:
-
- ValueError
-
If
lchdoes not have at least 3 channels (i.e., L*, C, and h).
See also
Notes
The h channel (i.e., hue) is expressed as an angle in range
(0, 2*pi).References
Examples
>>> from skimage import data >>> from skimage.color import rgb2lab, lch2lab, lab2lch >>> img = data.astronaut() >>> img_lab = rgb2lab(img) >>> img_lch = lab2lch(img_lab) >>> img_lab2 = lch2lab(img_lch)
-
skimage.color.luv2rgb(luv, *, channel_axis=-1)[source] -
Luv to RGB color space conversion.
- Parameters:
-
-
luv(…, C=3, …) array_like -
The image in CIE Luv format. By default, the final dimension denotes channels.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in RGB format. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If
luvis not at least 2-D with shape (…, C=3, …).
Notes
This function uses luv2xyz and xyz2rgb.
-
skimage.color.luv2xyz(luv, illuminant='D65', observer='2', *, channel_axis=-1)[source] -
CIE-Luv to XYZ color space conversion.
- Parameters:
-
-
luv(…, C=3, …) array_like -
The image in CIE-Luv format. By default, the final dimension denotes channels.
-
illuminant{“A”, “B”, “C”, “D50”, “D55”, “D65”, “D75”, “E”}, optional -
The name of the illuminant (the function is NOT case sensitive).
-
observer{“2”, “10”, “R”}, optional -
The aperture angle of the observer.
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in XYZ format. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If
luvis not at least 2-D with shape (…, C=3, …). - ValueError
-
If either the illuminant or the observer angle are not supported or unknown.
Notes
XYZ conversion weights use observer=2A. Reference whitepoint for D65 Illuminant, with XYZ tristimulus values of
(95.047, 100., 108.883). See functionxyz_tristimulus_values()for a list of supported illuminants.References
-
skimage.color.rgb2gray(rgb, *, channel_axis=-1)[source] -
Compute luminance of an RGB image.
- Parameters:
-
-
rgb(…, C=3, …) array_like -
The image in RGB format. By default, the final dimension denotes channels.
-
- Returns:
-
-
outndarray -
The luminance image - an array which is the same size as the input array, but with the channel dimension removed.
-
- Raises:
-
- ValueError
-
If
rgbis not at least 2-D with shape (…, C=3, …).
Notes
The weights used in this conversion are calibrated for contemporary CRT phosphors:
Y = 0.2125 R + 0.7154 G + 0.0721 B
If there is an alpha channel present, it is ignored.
References
Examples
>>> from skimage.color import rgb2gray >>> from skimage import data >>> img = data.astronaut() >>> img_gray = rgb2gray(img)
Using Polar and Log-Polar Transformations for Registration
Using Polar and Log-Polar Transformations for RegistrationRemoving small objects in grayscale images with a top hat filter
Removing small objects in grayscale images with a top hat filterFull tutorial on calibrating Denoisers Using J-Invariance
Full tutorial on calibrating Denoisers Using J-InvarianceGabors / Primary Visual Cortex “Simple Cells” from an Image
Gabors / Primary Visual Cortex "Simple Cells" from an Image
Region Boundary based Region adjacency graphs (RAGs)
Region Boundary based Region adjacency graphs (RAGs)
Comparison of segmentation and superpixel algorithms
Comparison of segmentation and superpixel algorithmsUse pixel graphs to find an object’s geodesic center
Use pixel graphs to find an object's geodesic center
-
skimage.color.rgb2hed(rgb, *, channel_axis=-1)[source] -
RGB to Haematoxylin-Eosin-DAB (HED) color space conversion.
- Parameters:
-
-
rgb(…, C=3, …) array_like -
The image in RGB format. By default, the final dimension denotes channels.
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in HED format. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If
rgbis not at least 2-D with shape (…, C=3, …).
References
[1]A. C. Ruifrok and D. A. Johnston, “Quantification of histochemical staining by color deconvolution.,” Analytical and quantitative cytology and histology / the International Academy of Cytology [and] American Society of Cytology, vol. 23, no. 4, pp. 291-9, Aug. 2001.
Examples
>>> from skimage import data >>> from skimage.color import rgb2hed >>> ihc = data.immunohistochemistry() >>> ihc_hed = rgb2hed(ihc)
-
skimage.color.rgb2hsv(rgb, *, channel_axis=-1)[source] -
RGB to HSV color space conversion.
- Parameters:
-
-
rgb(…, C=3, …) array_like -
The image in RGB format. By default, the final dimension denotes channels.
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in HSV format. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If
rgbis not at least 2-D with shape (…, C=3, …).
Notes
Conversion between RGB and HSV color spaces results in some loss of precision, due to integer arithmetic and rounding [1].
References
Examples
>>> from skimage import color >>> from skimage import data >>> img = data.astronaut() >>> img_hsv = color.rgb2hsv(img)
-
skimage.color.rgb2lab(rgb, illuminant='D65', observer='2', *, channel_axis=-1)[source] -
Conversion from the sRGB color space (IEC 61966-2-1:1999) to the CIE Lab colorspace under the given illuminant and observer.
- Parameters:
-
-
rgb(…, C=3, …) array_like -
The image in RGB format. By default, the final dimension denotes channels.
-
illuminant{“A”, “B”, “C”, “D50”, “D55”, “D65”, “D75”, “E”}, optional -
The name of the illuminant (the function is NOT case sensitive).
-
observer{“2”, “10”, “R”}, optional -
The aperture angle of the observer.
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in Lab format. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If
rgbis not at least 2-D with shape (…, C=3, …).
Notes
RGB is a device-dependent color space so, if you use this function, be sure that the image you are analyzing has been mapped to the sRGB color space.
This function uses rgb2xyz and xyz2lab. By default Observer=”2”, Illuminant=”D65”. CIE XYZ tristimulus values x_ref=95.047, y_ref=100., z_ref=108.883. See function
xyz_tristimulus_values()for a list of supported illuminants.References
-
skimage.color.rgb2luv(rgb, *, channel_axis=-1)[source] -
RGB to CIE-Luv color space conversion.
- Parameters:
-
-
rgb(…, C=3, …) array_like -
The image in RGB format. By default, the final dimension denotes channels.
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in CIE Luv format. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If
rgbis not at least 2-D with shape (…, C=3, …).
Notes
This function uses rgb2xyz and xyz2luv.
References
-
skimage.color.rgb2rgbcie(rgb, *, channel_axis=-1)[source] -
RGB to RGB CIE color space conversion.
- Parameters:
-
-
rgb(…, C=3, …) array_like -
The image in RGB format. By default, the final dimension denotes channels.
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in RGB CIE format. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If
rgbis not at least 2-D with shape (…, C=3, …).
References
Examples
>>> from skimage import data >>> from skimage.color import rgb2rgbcie >>> img = data.astronaut() >>> img_rgbcie = rgb2rgbcie(img)
-
skimage.color.rgb2xyz(rgb, *, channel_axis=-1)[source] -
RGB to XYZ color space conversion.
- Parameters:
-
-
rgb(…, C=3, …) array_like -
The image in RGB format. By default, the final dimension denotes channels.
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in XYZ format. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If
rgbis not at least 2-D with shape (…, C=3, …).
Notes
The CIE XYZ color space is derived from the CIE RGB color space. Note however that this function converts from sRGB.
References
Examples
>>> from skimage import data >>> img = data.astronaut() >>> img_xyz = rgb2xyz(img)
-
skimage.color.rgb2ycbcr(rgb, *, channel_axis=-1)[source] -
RGB to YCbCr color space conversion.
- Parameters:
-
-
rgb(…, C=3, …) array_like -
The image in RGB format. By default, the final dimension denotes channels.
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in YCbCr format. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If
rgbis not at least 2-D with shape (…, C=3, …).
Notes
Y is between 16 and 235. This is the color space commonly used by video codecs; it is sometimes incorrectly called “YUV”.
References
-
skimage.color.rgb2ydbdr(rgb, *, channel_axis=-1)[source] -
RGB to YDbDr color space conversion.
- Parameters:
-
-
rgb(…, C=3, …) array_like -
The image in RGB format. By default, the final dimension denotes channels.
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in YDbDr format. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If
rgbis not at least 2-D with shape (…, C=3, …).
Notes
This is the color space commonly used by video codecs. It is also the reversible color transform in JPEG2000.
References
-
skimage.color.rgb2yiq(rgb, *, channel_axis=-1)[source] -
RGB to YIQ color space conversion.
- Parameters:
-
-
rgb(…, C=3, …) array_like -
The image in RGB format. By default, the final dimension denotes channels.
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in YIQ format. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If
rgbis not at least 2-D with shape (…, C=3, …).
-
skimage.color.rgb2ypbpr(rgb, *, channel_axis=-1)[source] -
RGB to YPbPr color space conversion.
- Parameters:
-
-
rgb(…, C=3, …) array_like -
The image in RGB format. By default, the final dimension denotes channels.
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in YPbPr format. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If
rgbis not at least 2-D with shape (…, C=3, …).
References
-
skimage.color.rgb2yuv(rgb, *, channel_axis=-1)[source] -
RGB to YUV color space conversion.
- Parameters:
-
-
rgb(…, C=3, …) array_like -
The image in RGB format. By default, the final dimension denotes channels.
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in YUV format. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If
rgbis not at least 2-D with shape (…, C=3, …).
Notes
Y is between 0 and 1. Use YCbCr instead of YUV for the color space commonly used by video codecs, where Y ranges from 16 to 235.
References
-
skimage.color.rgba2rgb(rgba, background=(1, 1, 1), *, channel_axis=-1)[source] -
RGBA to RGB conversion using alpha blending [1].
- Parameters:
-
-
rgba(…, C=4, …) array_like -
The image in RGBA format. By default, the final dimension denotes channels.
-
backgroundarray_like -
The color of the background to blend the image with (3 floats between 0 to 1 - the RGB value of the background).
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in RGB format. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If
rgbais not at least 2D with shape (…, 4, …).
References
Examples
>>> from skimage import color >>> from skimage import data >>> img_rgba = data.logo() >>> img_rgb = color.rgba2rgb(img_rgba)
-
skimage.color.rgbcie2rgb(rgbcie, *, channel_axis=-1)[source] -
RGB CIE to RGB color space conversion.
- Parameters:
-
-
rgbcie(…, C=3, …) array_like -
The image in RGB CIE format. By default, the final dimension denotes channels.
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in RGB format. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If
rgbcieis not at least 2-D with shape (…, C=3, …).
References
Examples
>>> from skimage import data >>> from skimage.color import rgb2rgbcie, rgbcie2rgb >>> img = data.astronaut() >>> img_rgbcie = rgb2rgbcie(img) >>> img_rgb = rgbcie2rgb(img_rgbcie)
-
skimage.color.separate_stains(rgb, conv_matrix, *, channel_axis=-1)[source] -
RGB to stain color space conversion.
- Parameters:
-
-
rgb(…, C=3, …) array_like -
The image in RGB format. By default, the final dimension denotes channels.
- conv_matrix: ndarray
-
The stain separation matrix as described by G. Landini [1].
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in stain color space. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If
rgbis not at least 2-D with shape (…, C=3, …).
Notes
Stain separation matrices available in the
colormodule and their respective colorspace:-
hed_from_rgb: Hematoxylin + Eosin + DAB -
hdx_from_rgb: Hematoxylin + DAB -
fgx_from_rgb: Feulgen + Light Green -
bex_from_rgb: Giemsa stain : Methyl Blue + Eosin -
rbd_from_rgb: FastRed + FastBlue + DAB -
gdx_from_rgb: Methyl Green + DAB -
hax_from_rgb: Hematoxylin + AEC -
bro_from_rgb: Blue matrix Anilline Blue + Red matrix Azocarmine + Orange matrix Orange-G -
bpx_from_rgb: Methyl Blue + Ponceau Fuchsin -
ahx_from_rgb: Alcian Blue + Hematoxylin -
hpx_from_rgb: Hematoxylin + PAS
This implementation borrows some ideas from DIPlib [2], e.g. the compensation using a small value to avoid log artifacts when calculating the Beer-Lambert law.
References
[1][3]A. C. Ruifrok and D. A. Johnston, “Quantification of histochemical staining by color deconvolution,” Anal. Quant. Cytol. Histol., vol. 23, no. 4, pp. 291–299, Aug. 2001.
Examples
>>> from skimage import data >>> from skimage.color import separate_stains, hdx_from_rgb >>> ihc = data.immunohistochemistry() >>> ihc_hdx = separate_stains(ihc, hdx_from_rgb)
-
skimage.color.xyz2lab(xyz, illuminant='D65', observer='2', *, channel_axis=-1)[source] -
XYZ to CIE-LAB color space conversion.
- Parameters:
-
-
xyz(…, C=3, …) array_like -
The image in XYZ format. By default, the final dimension denotes channels.
-
illuminant{“A”, “B”, “C”, “D50”, “D55”, “D65”, “D75”, “E”}, optional -
The name of the illuminant (the function is NOT case sensitive).
-
observer{“2”, “10”, “R”}, optional -
One of: 2-degree observer, 10-degree observer, or ‘R’ observer as in R function grDevices::convertColor.
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in CIE-LAB format. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If
xyzis not at least 2-D with shape (…, C=3, …). - ValueError
-
If either the illuminant or the observer angle is unsupported or unknown.
Notes
By default Observer=”2”, Illuminant=”D65”. CIE XYZ tristimulus values x_ref=95.047, y_ref=100., z_ref=108.883. See function
xyz_tristimulus_values()for a list of supported illuminants.References
Examples
>>> from skimage import data >>> from skimage.color import rgb2xyz, xyz2lab >>> img = data.astronaut() >>> img_xyz = rgb2xyz(img) >>> img_lab = xyz2lab(img_xyz)
-
skimage.color.xyz2luv(xyz, illuminant='D65', observer='2', *, channel_axis=-1)[source] -
XYZ to CIE-Luv color space conversion.
- Parameters:
-
-
xyz(…, C=3, …) array_like -
The image in XYZ format. By default, the final dimension denotes channels.
-
illuminant{“A”, “B”, “C”, “D50”, “D55”, “D65”, “D75”, “E”}, optional -
The name of the illuminant (the function is NOT case sensitive).
-
observer{“2”, “10”, “R”}, optional -
The aperture angle of the observer.
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in CIE-Luv format. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If
xyzis not at least 2-D with shape (…, C=3, …). - ValueError
-
If either the illuminant or the observer angle are not supported or unknown.
Notes
By default XYZ conversion weights use observer=2A. Reference whitepoint for D65 Illuminant, with XYZ tristimulus values of
(95.047, 100., 108.883). See functionxyz_tristimulus_values()for a list of supported illuminants.References
Examples
>>> from skimage import data >>> from skimage.color import rgb2xyz, xyz2luv >>> img = data.astronaut() >>> img_xyz = rgb2xyz(img) >>> img_luv = xyz2luv(img_xyz)
-
skimage.color.xyz2rgb(xyz, *, channel_axis=-1)[source] -
XYZ to RGB color space conversion.
- Parameters:
-
-
xyz(…, C=3, …) array_like -
The image in XYZ format. By default, the final dimension denotes channels.
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in RGB format. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If
xyzis not at least 2-D with shape (…, C=3, …).
Notes
The CIE XYZ color space is derived from the CIE RGB color space. Note however that this function converts to sRGB.
References
Examples
>>> from skimage import data >>> from skimage.color import rgb2xyz, xyz2rgb >>> img = data.astronaut() >>> img_xyz = rgb2xyz(img) >>> img_rgb = xyz2rgb(img_xyz)
-
skimage.color.xyz_tristimulus_values(*, illuminant, observer, dtype=<class 'float'>)[source] -
Get the CIE XYZ tristimulus values.
Given an illuminant and observer, this function returns the CIE XYZ tristimulus values [2] scaled such that \(Y = 1\).
- Parameters:
-
-
illuminant{“A”, “B”, “C”, “D50”, “D55”, “D65”, “D75”, “E”} -
The name of the illuminant (the function is NOT case sensitive).
-
observer{“2”, “10”, “R”} -
One of: 2-degree observer, 10-degree observer, or ‘R’ observer as in R function
grDevices::convertColor[3]. - dtype: dtype, optional
-
Output data type.
-
- Returns:
-
-
valuesarray -
Array with 3 elements \(X, Y, Z\) containing the CIE XYZ tristimulus values of the given illuminant.
-
- Raises:
-
- ValueError
-
If either the illuminant or the observer angle are not supported or unknown.
Notes
The CIE XYZ tristimulus values are calculated from \(x, y\) [1], using the formula
\[X = x / y\]\[Y = 1\]\[Z = (1 - x - y) / y\]The only exception is the illuminant “D65” with aperture angle 2° for backward-compatibility reasons.
References
Examples
Get the CIE XYZ tristimulus values for a “D65” illuminant for a 10 degree field of view
>>> xyz_tristimulus_values(illuminant="D65", observer="10") array([0.94809668, 1. , 1.07305136])
-
skimage.color.ycbcr2rgb(ycbcr, *, channel_axis=-1)[source] -
YCbCr to RGB color space conversion.
- Parameters:
-
-
ycbcr(…, C=3, …) array_like -
The image in YCbCr format. By default, the final dimension denotes channels.
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in RGB format. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If
ycbcris not at least 2-D with shape (…, C=3, …).
Notes
Y is between 16 and 235. This is the color space commonly used by video codecs; it is sometimes incorrectly called “YUV”.
References
-
skimage.color.ydbdr2rgb(ydbdr, *, channel_axis=-1)[source] -
YDbDr to RGB color space conversion.
- Parameters:
-
-
ydbdr(…, C=3, …) array_like -
The image in YDbDr format. By default, the final dimension denotes channels.
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in RGB format. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If
ydbdris not at least 2-D with shape (…, C=3, …).
Notes
This is the color space commonly used by video codecs, also called the reversible color transform in JPEG2000.
References
-
skimage.color.yiq2rgb(yiq, *, channel_axis=-1)[source] -
YIQ to RGB color space conversion.
- Parameters:
-
-
yiq(…, C=3, …) array_like -
The image in YIQ format. By default, the final dimension denotes channels.
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in RGB format. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If
yiqis not at least 2-D with shape (…, C=3, …).
-
skimage.color.ypbpr2rgb(ypbpr, *, channel_axis=-1)[source] -
YPbPr to RGB color space conversion.
- Parameters:
-
-
ypbpr(…, C=3, …) array_like -
The image in YPbPr format. By default, the final dimension denotes channels.
-
channel_axisint, optional -
This parameter indicates which axis of the array corresponds to channels.
Added in version 0.19:
channel_axiswas added in 0.19.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in RGB format. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If
ypbpris not at least 2-D with shape (…, C=3, …).
References
-
skimage.color.yuv2rgb(yuv, *, channel_axis=-1)[source] -
YUV to RGB color space conversion.
- Parameters:
-
-
yuv(…, C=3, …) array_like -
The image in YUV format. By default, the final dimension denotes channels.
-
- Returns:
-
-
out(…, C=3, …) ndarray -
The image in RGB format. Same dimensions as input.
-
- Raises:
-
- ValueError
-
If
yuvis not at least 2-D with shape (…, C=3, …).
References
© 2019 the scikit-image team
Licensed under the BSD 3-clause License.
https://scikit-image.org/docs/0.25.x/api/skimage.color.html